Protein Evolution and Post-translational Modifications
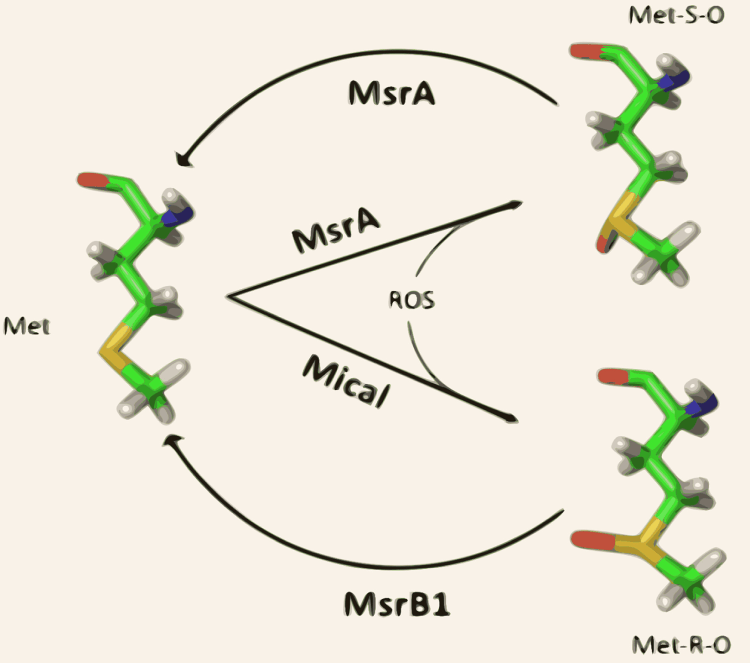
Our research interest spans various aspects of computational evolutionary biology. Generally speaking, our work seeks to better understand how proteins evolve [1]. In particular, we are interested in understanding the very diverse roles that methionine residues fulfil in proteins (reviewed in [2] and [3]), and how these functions have evolved in organisms. Along these lines, we have devoted great attention to methionine sulfoxidation as a bone-fide post-translational modification [3-7].
We tackle these problems by gathering and curating experimental data and developing computational tools to be applied on the gathered data. In this sense, we have developed and made available a software package, ptm, for the analysis of post-translational modifications [8], as well as a collection of data and references to methionines that may be involved in regulatory processes. Thus, MetOSite is a web-based searchable database that currently contains over 7500 methionine sulfoxide sites found in over 3700 different proteins from more than 30 species [9].
Selected References
- Multisite phosphorylation provides a reliable mechanism for making decisions in noisy environments. FEBS J. 2018 Oct;285(20):3729-3737. doi: 10.1111/febs.14636. Epub 2018 Sep 4.
- Methionine in proteins: The Cinderella of the proteinogenic amino acids. Protein Sci. 2019 Oct;28(10):1785-1796. doi: 10.1002/pro.3698.
- The role of methionine residues in the regulation of liquid-liquid phase separation. Biomolecules. 2021 Aug 21;11(8):1248. doi: 10.3390/biom11081248.
- Susceptibility of protein methionine oxidation in response to hydrogen peroxide treatment –ex vivo versus in vitro-: a computational insight. Antioxidants (Basel). 2020 Oct 13;9(10):987. doi: 10.3390/antiox9100987.
- Inferring methionine sulfoxidation and serine phosphorylation crosstalk from phylogenetic analyses. BMC Evol Biol. 2017 Jul 27;17(1):171. doi: 10.1186/s12862-017-1017-9.
- Methionine residues around phosphorylation sites are preferentially oxidized in vivo under stress conditions. Sci Rep. 2017 Jan 12;7:40403. doi: 10.1038/srep40403.
- Sulfur atoms from methionines interacting with aromatic residues are less prone to oxidation. Sci Rep. 2015 Nov 24;5:16955. doi: 10.1038/srep16955.
- ptm: an R package for the study of methionine sulfoxidation and other post-translational modifications. Bioinformatics. 2021 May 8:btab348. doi: 10.1093/bioinformatics/btab348.
- MetOSite: an integrated resource for the study of methionine residue sulfoxidation. Bioinformatics. 2019 Nov 1;35(22):4849-4850. doi: 10.1093/bioinformatics/btz462